import numpy as np
n_groups = 1000
anchors = np.random.randn(n_groups, 3) * 1 # spread out in 3D
target = np.random.randint(0, 3, n_groups)
seconds = anchors + np.random.randn(n_groups, 3) * 0.3 # close to anchor
thirds = anchors + np.random.randn(n_groups, 3) * 0.3 # close to anchor
sets_list = [anchors, seconds, thirds]
coords = np.vstack(sets_list) # stacked: [A|B|C] along batch dim
fig = plot_embeddings_3d(coords, color_by='triplets' if len(sets_list)==3 else 'pairs', title='Triplet test', target=target)
fig.show()viz
UMAP
cpu_umap_project
def cpu_umap_project(
embeddings, n_components:int=3, n_neighbors:int=15, min_dist:float=0.1, random_state:int=42
):
Project embeddings to n_components dimensions via UMAP (on CPU)
cuml_umap_project
def cuml_umap_project(
embeddings, n_components:int=3, n_neighbors:int=15, min_dist:float=0.1, random_state:int=42
):
Project embeddings to n_components dimensions via cuML UMAP (GPU)
umap_project
def umap_project(
embeddings, kwargs:VAR_KEYWORD
):
Calls one of two preceding UMAP routines based on device availability.
PCA
cuml_pca_project
def cuml_pca_project(
embeddings, n_components:int=3
):
Project embeddings to n_components dimensions via cuML PCA (GPU)
cpu_pca_project
def cpu_pca_project(
embeddings, n_components:int=3
):
Project embeddings to n_components dimensions via sklearn PCA (CPU)
pca_project
def pca_project(
embeddings, kwargs:VAR_KEYWORD
):
Calls GPU or CPU PCA based on availability
3D Plotly Scatterplots
plot_embeddings_3d
def plot_embeddings_3d(
coords, color_by:str='pairs', file_idx:NoneType=None, deltas:NoneType=None, title:str='Embeddings',
target:NoneType=None, debug:bool=False
):
3D scatter plot of embeddings. color_by: ‘none’, ‘file’, ‘pairs’, or ‘triplets’
Test code for triplet viz:
Main Routine
Calls the preceding routines
Testing _subsample:
n_pairs, dim = 5, 1 # data points
z1 = 200*torch.arange(n_pairs).unsqueeze(-1).unsqueeze(-1)
z2 = z1 + 1
zs = torch.cat([z1, z2], dim=0).view(-1, dim)
print("zs.shape = ",zs.shape)
indices = torch.arange(2*n_pairs)
deltas = torch.randint(0,12,(2*n_pairs, 2))
print("zs = \n",zs)
print("indices =",indices)
data_perm, indices2, deltas2, _ = _subsample(zs, indices, deltas, max_points=2*(n_pairs-2), debug=True)
print("data_perm.shape = ",data_perm.shape,", data_perm = \n",data_perm)zs.shape = torch.Size([10, 1])
zs =
tensor([[ 0],
[200],
[400],
[600],
[800],
[ 1],
[201],
[401],
[601],
[801]])
indices = tensor([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
data_perm.shape = torch.Size([6, 1]) , data_perm =
tensor([[600],
[200],
[800],
[601],
[201],
[801]])Ok. That does what I expect. Moving on…
make_emb_viz
def make_emb_viz(
enc_outs, epoch:int=-1, encoder:NoneType=None, batch:NoneType=None, title:str='Embeddings', max_points:int=5000,
do_umap:bool=False, debug:bool=False, color_by:NoneType=None
):
this is the main viz routine, showing different groups of embeddings
Testing visualization:
import plotly.io as pio
pio.renderers.default = 'notebook'
from midi_rae.core import PatchState, HierarchicalPatchState, EncoderOutput
bs = 32 # Batch size = 32
# Mimic Swin hierarchy: (grid_size, embed_dim) — coarsest to finest
level_specs = [(1,256), (2,128), (4,64), (8,32), (16,16), (32,8)]
def make_fake_enc_out(other=None, noise=0.1):
levels = []
for grid, dim in level_specs:
N = grid * grid
pos = torch.stack([torch.tensor([r, c]) for r in range(grid) for c in range(grid)])
emb = other_levels[len(levels)].emb + noise * torch.randn(bs, N, dim) if other else torch.randn(bs, N, dim)
ne = torch.ones(bs, N, dtype=torch.bool)
if grid >= 8: ne = torch.rand(bs, N) > 0.5 # make some empty at finer levels
levels.append(PatchState(emb=emb, pos=pos, non_empty=ne, mae_mask=torch.ones(N, dtype=torch.bool)))
patches = HierarchicalPatchState(levels=levels)
return EncoderOutput(
patches=patches,
full_pos=torch.cat([l.pos for l in levels]),
full_non_empty=torch.cat([l.non_empty for l in levels], dim=1),
mae_mask=torch.cat([l.mae_mask for l in levels]),
)
enc_out1 = make_fake_enc_out()
other_levels = enc_out1.patches.levels # so enc_out2 is a noisy copy
enc_out2 = make_fake_enc_out(other=True)
enc_out3 = make_fake_enc_out(other=True)
batch = {'file_idx': torch.arange(bs), 'deltas': torch.randint(0, 12, (bs, 2)), 'target': torch.randint(0,3,(bs,))}
figs = make_emb_viz([enc_out1, enc_out2, enc_out3], title='testing', batch=batch, do_umap=False, debug=True)
figs[5]['joint_non_empty']['pca'].show()We should make some code to display the various levels neatly in a table:
show_fig_table
def show_fig_table(
figs
):
Display all PCA figs in a grid: rows=levels, cols=non_empty|empty
show_fig_table(figs)Reconstructions
expand_patch_mask
def expand_patch_mask(
mae_mask, grid_h, grid_w, patch_size
):
Expand patch-level mask (N,) to pixel-level mask (H, W)
do_recon_eval
def do_recon_eval(
recon, real, mae_mask:NoneType=None, patch_size:int=16, eps:float=1e-08, return_maps:bool=False
):
Evaluate recon accuracy, optionally only on masked patches
patches_to_img
def patches_to_img(
recon_patches, img_real, patch_size:int=16, mae_mask:NoneType=None
):
Convert image patches to full image. Copy over real patches where appropriate.
viz_mae_recon
def viz_mae_recon(
recon, img_real, enc_out:NoneType=None, epoch:int=-1, patch_size:int=16, debug:bool=False,
return_maps:bool=False
):
Show how our LightweightMAEDecoder is doing (during encoder training)
Testing code:
from midi_rae.core import *
import matplotlib.pyplot as plt
B, patch_size = 4, 16
img_real = (torch.rand(B, 1, 128, 128) > 0.7).float() # fake sparse piano roll
recon = torch.randn(B, 65, patch_size**2) # 64 patches + CLS, raw logits
mae_mask = torch.ones(65, dtype=torch.bool)
mae_mask[1::2] = False # mask every other patch (skip CLS at 0)
enc_out = EncoderOutput(
patches=HierarchicalPatchState(levels=[
PatchState(emb=torch.randn(B,1,256), pos=torch.tensor([[-1,-1]]), non_empty=torch.ones(B,1,dtype=torch.bool), mae_mask=mae_mask[0:1]),
PatchState(emb=torch.randn(B,64,256), pos=torch.zeros(64,2), non_empty=torch.ones(B,64,dtype=torch.bool), mae_mask=mae_mask[1:]),
]),
full_pos=torch.zeros(65,2), full_non_empty=torch.ones(B,65,dtype=torch.bool), mae_mask=mae_mask,
)
grid_recon, grid_real, grid_map, evals = viz_mae_recon(recon, img_real, enc_out=enc_out, epoch=0, debug=True, return_maps=True)
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, figsize=(12, 6))
ax1.imshow(grid_real.permute(1,2,0), cmap='gray'); ax1.set_title('Real')
ax2.imshow(grid_recon.permute(1,2,0), cmap='gray'); ax2.set_title('Recon')
ax3.imshow(grid_map.permute(1,2,0)); ax3.set_title('Map')
plt.show()
print(', '.join(f"{k}: {v.item():.4f}" for k, v in evals.items() if not k.endswith('map')))mae_mask: shape=torch.Size([64]), pct_visible=0.500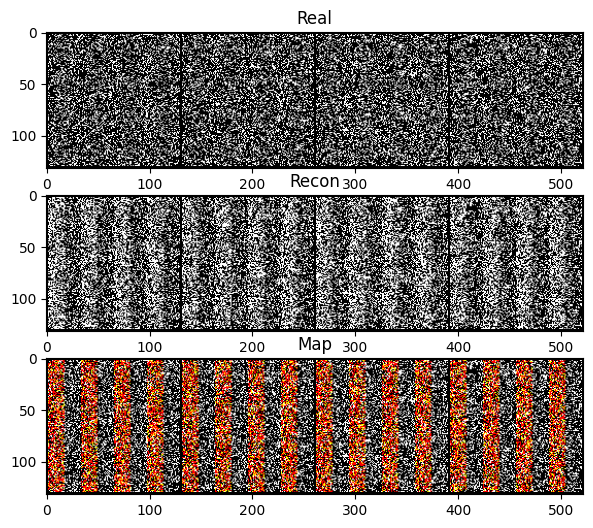
precision: 0.9141, recall: 0.4991, specificity: 0.4996, f1: 0.6456