import os
assert False == os.path.isdir('/app/data'), "Do not try to run this on solveit. The memory requirements will crash the VM."Inspect
(Notebooke only) Interactive exploration of trained encoder and embeddings
import torch
import torch.nn.functional as F
from torch.utils.data import DataLoader
from omegaconf import OmegaConf
import matplotlib.pyplot as plt
from PIL import Image
from IPython.display import display
import plotly.io as pio
pio.renderers.default = 'notebook'
from tqdm.auto import tqdm
from midi_rae.data import PRPairDataset, ShiftedTripletDataset, TARGET_NAMES, SCHEME_NAMES
from midi_rae.vit import ViTEncoder, ViTDecoder
from midi_rae.swin import SwinEncoder, SwinDecoder
from midi_rae.utils import load_checkpoint
from midi_rae.viz import make_emb_viz, viz_mae_recon, show_fig_tableConfig
#cfg = OmegaConf.load('../configs/config.yaml')
cfg = OmegaConf.load('../configs/config_swin.yaml')
#device = 'cuda' if torch.cuda.is_available() else 'mps' if torch.backends.mps.is_available() else 'cpu'
device = 'cpu' # leave GPU free for training while we do analysis here.
print(f'device = {device}')device = cpuLoad Dataset
#val_ds = PRPairDataset(image_dataset_dir=cfg.data.path, split='val', max_shift_x=cfg.training.max_shift_x, max_shift_y=cfg.training.max_shift_y)
val_ds = ShiftedTripletDataset(image_dataset_dir=cfg.data.path, split='val', max_shift_x=cfg.training.max_shift_x, max_shift_y=cfg.training.max_shift_y)
val_dl = DataLoader(val_ds, batch_size=cfg.training.batch_size, num_workers=4, drop_last=True)
print(f'Loaded {len(val_ds)} validation samples, batch_size = {cfg.training.batch_size}')Loading 91 val files from /home/shawley/datasets/POP909_images_basic... Finished loading.
Loaded 9100 validation samples, batch_size = 370Inspect Data
batch = next(iter(val_dl))
img1, img2, deltas, file_idx = batch['img1'].to(device), batch['img2'].to(device), batch['deltas'].to(device), batch['file_idx'].to(device)
print("img1.shape, deltas.shape, file_idx.shape =",tuple(img1.shape), tuple(deltas.shape), tuple(file_idx.shape))img1.shape, deltas.shape, file_idx.shape = (370, 1, 128, 128) (370, 2, 2) (370,)# Show a sample image pair
idx = 0
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
axes[0].imshow(img1[idx, 0].cpu(), cmap='gray')
axes[0].set_title(f'Image 1 (file_idx={file_idx[idx].item()})')
axes[1].imshow(img2[idx, 0].cpu(), cmap='gray')
axes[1].set_title(f'Image 2 (deltas = {deltas[idx].cpu().int().numpy()})')
plt.tight_layout()
plt.show()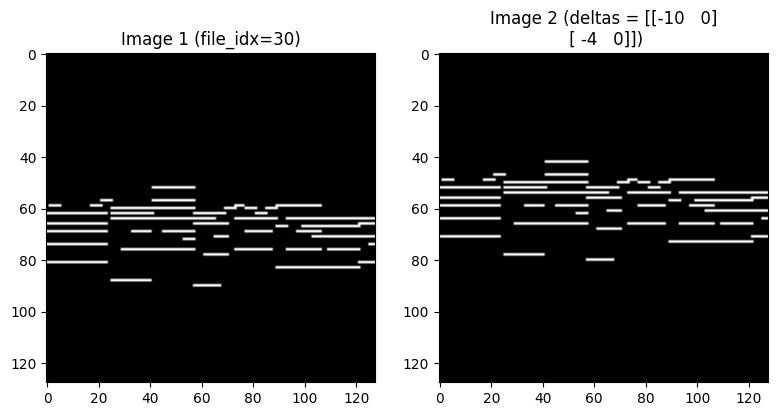
Load Encoder from Checkpoint
if cfg.model.get('encoder', 'vit') == 'swin':
encoder = SwinEncoder(img_height=cfg.data.image_size, img_width=cfg.data.image_size,
patch_h=cfg.model.patch_h, patch_w=cfg.model.patch_w,
embed_dim=cfg.model.embed_dim, depths=cfg.model.depths,
num_heads=cfg.model.num_heads, window_size=cfg.model.window_size,
mlp_ratio=cfg.model.mlp_ratio, drop_path_rate=cfg.model.drop_path_rate).to(device)
else:
encoder = ViTEncoder(cfg.data.in_channels, cfg.data.image_size, cfg.model.patch_size,
cfg.model.dim, cfg.model.depth, cfg.model.heads).to(device)
#encoder = load_checkpoint(encoder, cfg.get('encoder_ckpt', f'../checkpoints/{encoder.__class__.__name__}__best.pt'))
encoder = load_checkpoint(encoder, '../checkpoints/SwinEncoder_NoSim_best.pt')
encoder.eval()
print(f"Loaded {encoder.__class__.__name__}")>>> Loaded SwinEncoder checkpoint from ../checkpoints/SwinEncoder_NoSim_best.pt
Loaded SwinEncoderRun Batch Through Encoder
with torch.no_grad():
with torch.autocast(device_type=device, dtype=torch.bfloat16):
enc_out1 = encoder(img1)
enc_out2 = encoder(img2)Visualize Embeddings
NOTE: This will visualize all embeddings in the entire batch, not just the single pair of images shown above.
figs = make_emb_viz((enc_out1, enc_out2), encoder=encoder, batch=batch, do_umap=False)
show_fig_table(figs)SVD Analysis
def svd_analysis(enc_out, level=1, title='', top_k=20):
"Run SVD on encoder output, plot singular value spectrum and cumulative variance"
z = enc_out.patches[level].emb.detach().cpu().float().reshape(-1, enc_out.patches[level].dim) # flatten batch
z = z - z.mean(dim=0) # center
U, S, Vt = torch.linalg.svd(z, full_matrices=False) # Vt for "V transpose" (technically it's "V hermitian" but we've got real data)
var_exp = (S**2) / (S**2).sum()
cum_var = var_exp.cumsum(0)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(18, 5))
ax1.semilogy(S.numpy()); ax1.axvline(x=top_k, color='r', ls='--', alpha=0.5)
ax1.set(xlabel='Component', ylabel='Singular value', title=f'{title} Singular Values')
ax2.bar(range(top_k), var_exp[:top_k].numpy())
ax2.set(xlabel='Component', ylabel='Variance explained', title=f'{title} Top {top_k} Variance')
ax3.plot(cum_var.numpy()); ax3.axhline(y=0.9, color='r', ls='--', alpha=0.5, label='90%')
ax3.set(xlabel='Component', ylabel='Cumulative variance', title=f'{title} Cumulative Variance')
ax3.legend()
plt.tight_layout(); plt.show()
n90 = (cum_var < 0.9).sum().item() + 1
print(f"{title}: {n90} components for 90% variance, top-1 explains {var_exp[0]:.1%}")
return S, U, Vt, var_expS, U, Vt, var_exp = svd_analysis(enc_out2, title='Finest Patches,')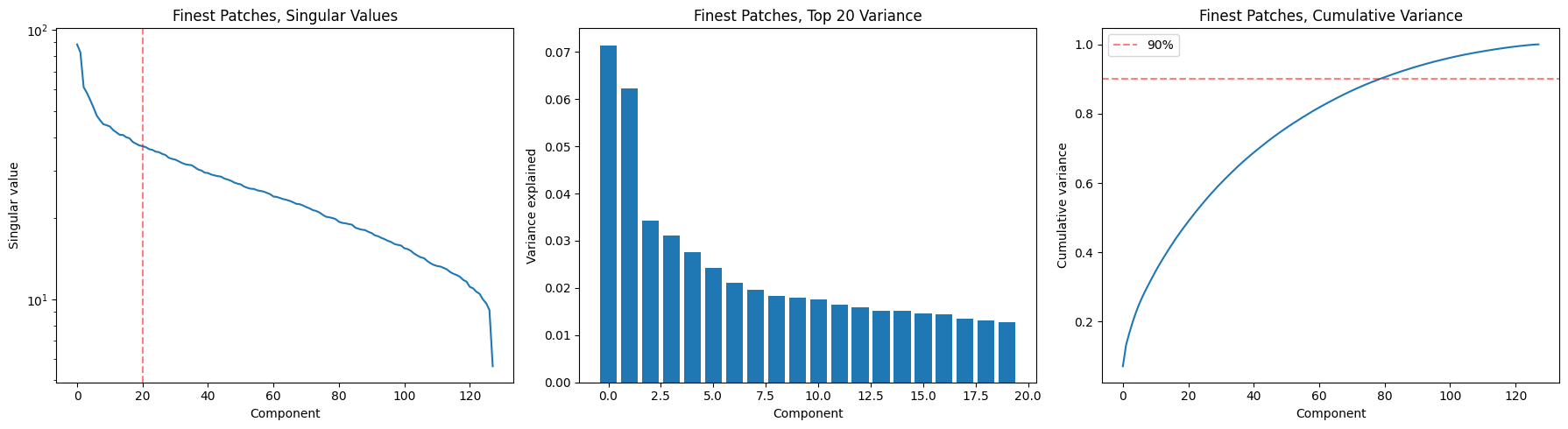
Finest Patches,: 80 components for 90% variance, top-1 explains 7.1%cls_S, cls_U, cls_Vt, cls_var_exp = svd_analysis(enc_out2, level=0, title='CLS / Coarsest Patch,')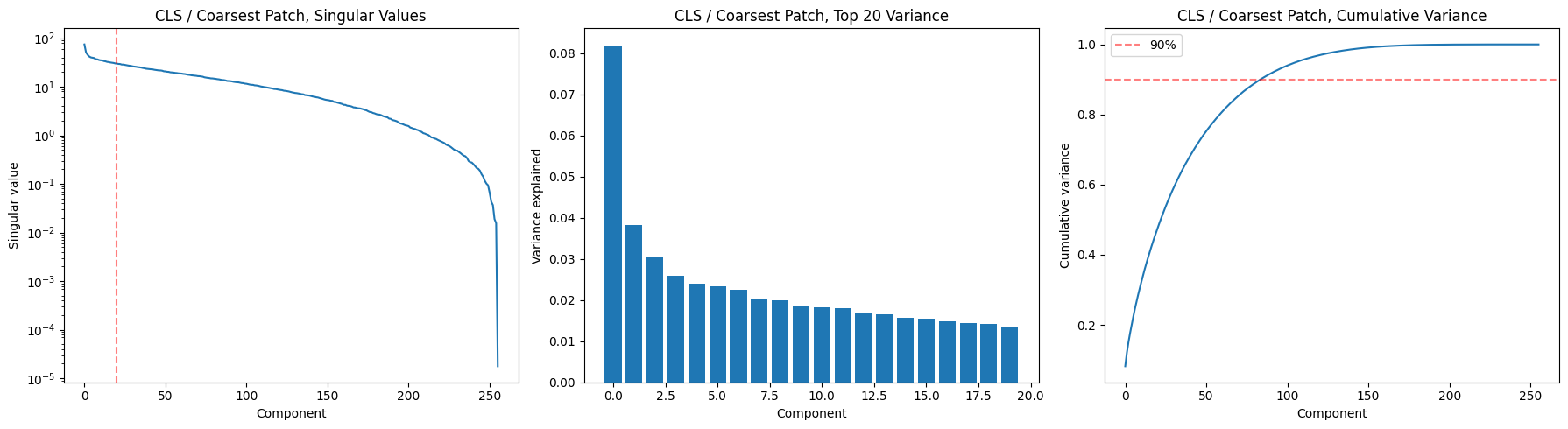
CLS / Coarsest Patch,: 85 components for 90% variance, top-1 explains 8.2%Two key takeaways:
- Patches need 178/256 dims for 90%. The representation is highly distributed with no dominant direction. This means the encoder is using nearly all its capacity, which is healthy (no dimensional collapse). But it also suggests rhythm and pitch aren’t cleanly factored — if they were, you’d expect a sharper elbow in the spectrum (the first 1 or 2 components notwithstanding).
- CLS only needs 23/256 dims. The global summary is much more compressed. That’s interesting for generation: it suggests the “gist” of a musical passage lives in a ~23-dimensional subspace. The gradual slope in the top-20 bars (no single dominant component) means it’s not collapsing to a trivial representation either.
Decoder Performance
if cfg.model.get('encoder', 'vit') == 'swin': # decoder should match encoder
decoder = SwinDecoder(img_height=cfg.data.image_size, img_width=cfg.data.image_size,
patch_h=cfg.model.patch_h, patch_w=cfg.model.patch_w,
embed_dim=cfg.model.embed_dim,
depths=cfg.model.get('dec_depths', cfg.model.depths),
num_heads=cfg.model.get('dec_num_heads', cfg.model.num_heads)).to(device)
else:
decoder = ViTDecoder(cfg.data.in_channels, (cfg.data.image_size, cfg.data.image_size),
cfg.model.patch_size, cfg.model.dim,
cfg.model.get('dec_depth', 4), cfg.model.get('dec_heads', 8)).to(device)
name = decoder.__class__.__name__
print("Name = ",name)
decoder = load_checkpoint(decoder, cfg.get('encoder_ckpt', f'../checkpoints/{decoder.__class__.__name__}__best.pt'))Name = SwinDecoder
>>> Loaded SwinDecoder checkpoint from ../checkpoints/SwinDecoder__best.ptwith torch.autocast(device_type='cpu', dtype=torch.bfloat16):
recon_logits = decoder(enc_out2)
img_recon = torch.sigmoid(recon_logits).float().cpu()
img_real = img2
print("img_recon.shape, img_real.shape =",img_recon.shape, img_real.shape)img_recon.shape, img_real.shape = torch.Size([256, 1, 128, 128]) torch.Size([256, 1, 128, 128])grid_recon, grid_real, grid_map, evals = viz_mae_recon(img_recon, img_real, enc_out=None, epoch=0, debug=False, return_maps=True)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(10, 5))
ax1.imshow(grid_real.permute(1,2,0), cmap='gray'); ax1.set_title('Real')
ax2.imshow(grid_recon.permute(1,2,0), cmap='gray'); ax2.set_title('Recon')
ax3.imshow(grid_map.permute(1,2,0)); ax3.set_title('Map')
plt.show()
print(', '.join(f"{k}: {v.item():.4f}" for k, v in evals.items() if not k.endswith('map')))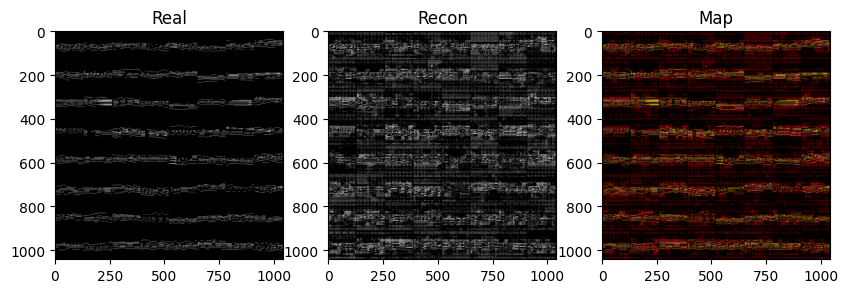
precision: 0.0978, recall: 0.3414, specificity: 0.8724, f1: 0.1521In the next cell we’re gonna plot an image showing the maps as a very large image, we’re gonna hide it from the LLM because it doesn’t need to see it and we wanna spare the context.
img = Image.fromarray((grid_map*255).permute(1,2,0).byte().numpy())
display(img)Pitch-time factorization
Cosine Histograms and PCA plots
batch_size = 256
ds = ShiftedTripletDataset(split='val')
subset = torch.utils.data.Subset(ds, range(batch_size * 10)) # 10 samples is enough
dl = DataLoader(subset, batch_size=batch_size, shuffle=False)
plt.close('all')
print("Collecting difference vectors...")
targets, schemes = [], []
for i, batch in enumerate(tqdm(dl)):
img1, img2, img3 = batch['img1'], batch['img2'], batch['img3']
deltas, scheme, target, scheme = batch['deltas'], batch['scheme'], batch['target'], batch['scheme']
targets.append(target)
schemes.append(scheme)
with torch.no_grad():
enc_out1 = encoder(img1) # anchor
enc_out2 = encoder(img2) # crop 1
enc_out3 = encoder(img3) # crop 2
z1 = [lvl.emb for lvl in enc_out1.patches.levels]
z2 = [lvl.emb for lvl in enc_out2.patches.levels]
z3 = [lvl.emb for lvl in enc_out3.patches.levels]
n_levels = enc_out1.patches.num_levels
# lazy-initialize the accumulation lists
if i == 0: d2_vecs, d3_vecs = [[] for _ in range(n_levels)], [[] for _ in range(n_levels)]
for lev in range(n_levels):
d2, d3 = z2[lev]-z1[lev], z3[lev]-z1[lev]
d2, d3 = (z2[lev]-z1[lev]).mean(dim=1), (z3[lev]-z1[lev]).mean(dim=1) # mean pooling across patches
d2_vecs[lev].append(d2)
d3_vecs[lev].append(d3)
del dl
targets = torch.cat(targets,dim=0)
schemes = torch.cat(schemes,dim=0)
# print("Performing per-level analysis: (takes a while)")
# OUR_TARGET_NAMES = {-1.0: 'anti-parallel', 0.0: 'orthogonal', 1.0: 'parallel'}
# for lev in range(n_levels):
# d2 = torch.cat(d2_vecs[lev], dim=0)
# d3 = torch.cat(d3_vecs[lev], dim=0)
# cos = F.cosine_similarity(d2, d3, dim=-1)
# print(f"Level = {lev}: d2 shape = {d2[lev].shape}, cos NaNs = {cos.isnan().sum()}")
# n = min(len(cos), 20000)
# idx = torch.randperm(len(cos))[:n]
# cos_sub, targets_sub = cos[idx], targets[idx]
# fig, ax = plt.subplots(1, 3, figsize=(8,2))
# for i, (t_val, name) in enumerate(OUR_TARGET_NAMES.items()):
# mask = (targets_sub - t_val).abs() < 1e-3
# ax[i].hist(cos_sub[mask].numpy().flatten(), bins=50, alpha=0.5, label=f"Level {lev}:\n N= {len(cos_sub[mask].numpy().flatten())}, {name}")
# ax[i].set_ylabel('count')
# ax[i].set_xlabel('cosine')
# ax[i].set_xlim(-1, 1)
# ax[i].legend(fancybox=True, framealpha=0.3)
# plt.show()
# plt.close()Loading 91 val files from ~/datasets/POP909_images_basic/... Finished loading.
Collecting difference vectors...from sklearn.decomposition import PCA
for lev in range(n_levels):
d2 = torch.cat(d2_vecs[lev], dim=0)
d3 = torch.cat(d3_vecs[lev], dim=0)
# Separate into pitch and time diffs by scheme
s0, s1, s2 = schemes == 0, schemes == 1, schemes == 2
pitch_d = torch.cat([d2[s0], d3[s0], d2[s2]], dim=0)
time_d = torch.cat([d2[s1], d3[s1], d3[s2]], dim=0)
all_diffs = torch.cat([pitch_d, time_d], dim=0).numpy()
pca = PCA(n_components=2).fit(all_diffs)
proj = pca.transform(all_diffs)
n_pitch = len(pitch_d)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 5))
fig.suptitle(f'Level {lev}', fontsize=14)
ax1.scatter(proj[:n_pitch, 0], proj[:n_pitch, 1], alpha=0.3, s=10, label='pitch Δ', c='blue')
ax1.scatter(proj[n_pitch:, 0], proj[n_pitch:, 1], alpha=0.3, s=10, label='time Δ', c='green')
ax1.set_xlabel('PC1'); ax1.set_ylabel('PC2'); ax1.legend()
ax1.set_title('PCA of difference vectors'); ax1.set_aspect('equal')
# Cross-type cosines
n_min = min(len(pitch_d), len(time_d))
cos_cross = F.cosine_similarity(pitch_d[:n_min], time_d[:n_min], dim=-1).numpy()
# Same-type cosines by target
same_type = s0 | s1
par = same_type & (targets > 0.5)
anti = same_type & (targets < -0.5)
cos_par = F.cosine_similarity(d2[par], d3[par], dim=-1).numpy()
cos_anti = F.cosine_similarity(d2[anti], d3[anti], dim=-1).numpy()
ax2.hist(cos_cross, bins=40, alpha=0.5, label='pitch vs time (want ≈0)', density=True)
ax2.hist(cos_par, bins=40, alpha=0.5, label='same-sign (want ≈+1)', density=True)
ax2.hist(cos_anti, bins=40, alpha=0.5, label='opp-sign (want ≈−1)', density=True)
ax2.set_xlabel('cosine similarity'); ax2.legend(); ax2.set_title('Cosine similarity distributions')
plt.tight_layout()
plt.show()
plt.close()
print(f"Level {lev}: cross={cos_cross.mean():.3f}, parallel={cos_par.mean():.3f}, anti={cos_anti.mean():.3f}\n")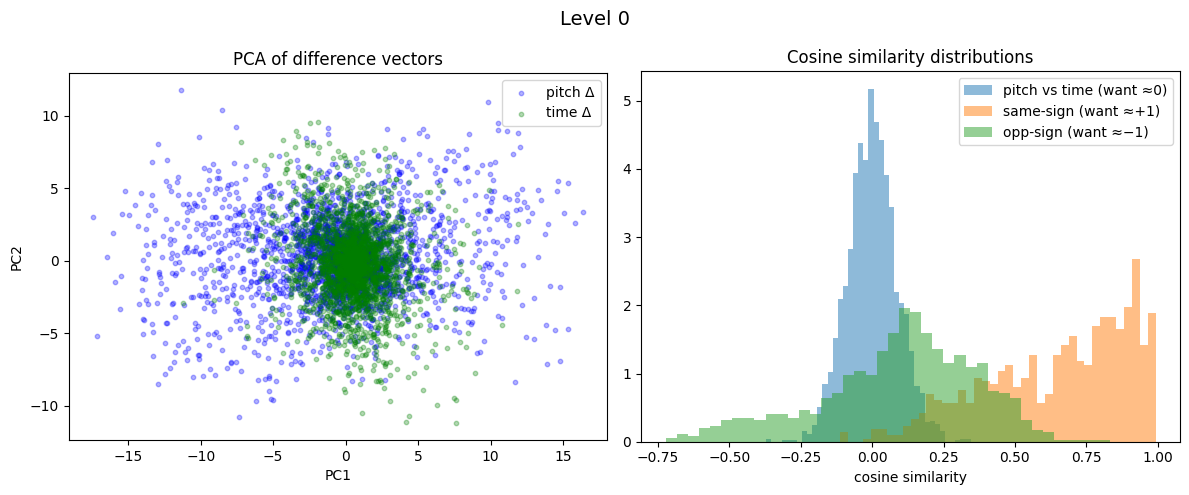
Level 0: cross=0.001, parallel=0.649, anti=0.092
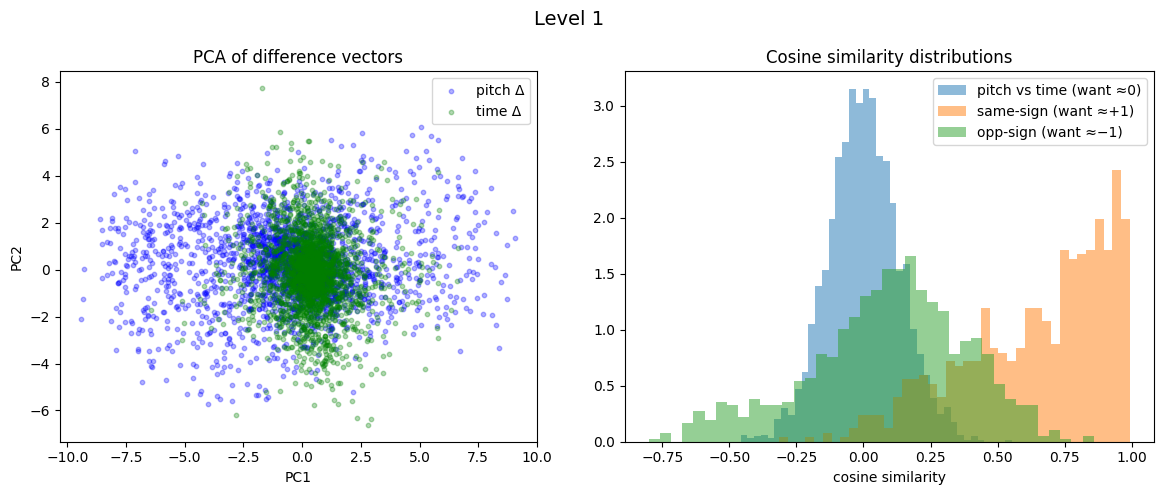
Level 1: cross=0.001, parallel=0.644, anti=0.089
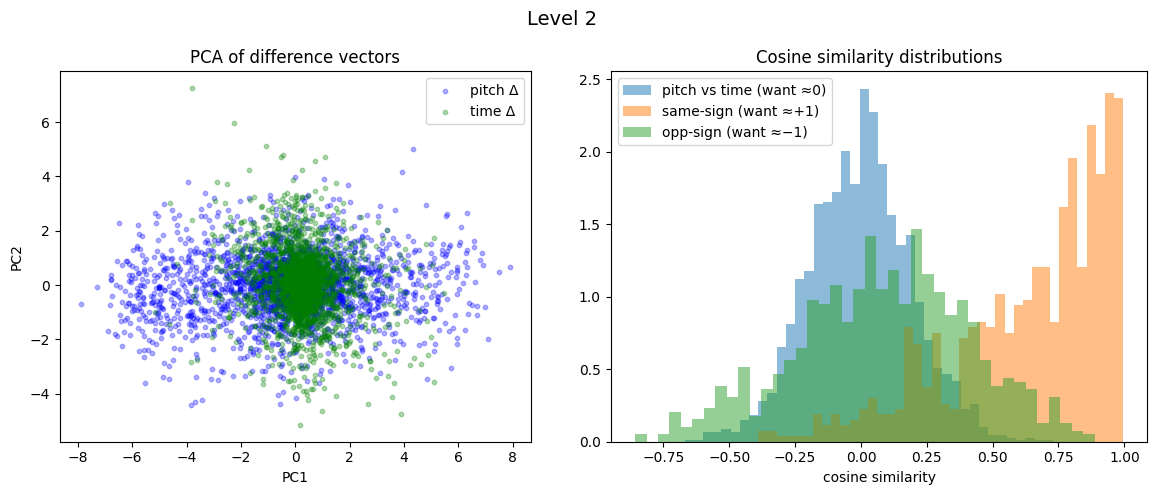
Level 2: cross=-0.003, parallel=0.639, anti=0.082
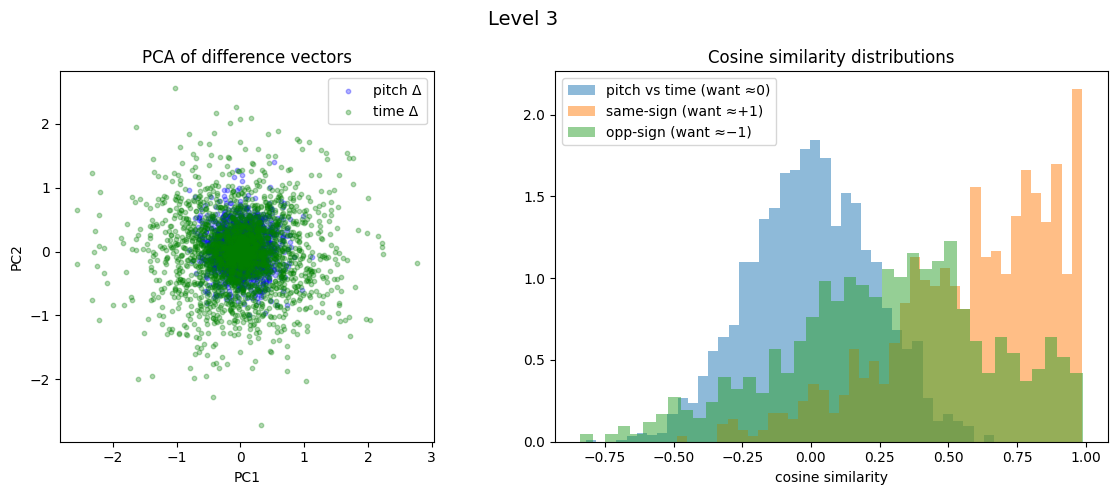
Level 3: cross=0.002, parallel=0.587, anti=0.280
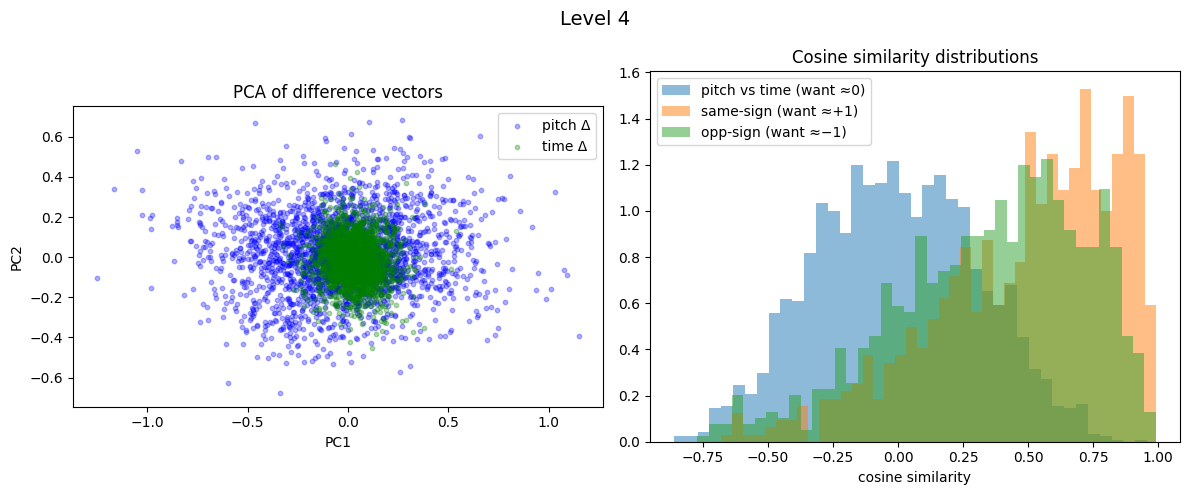
Level 4: cross=-0.005, parallel=0.491, anti=0.352
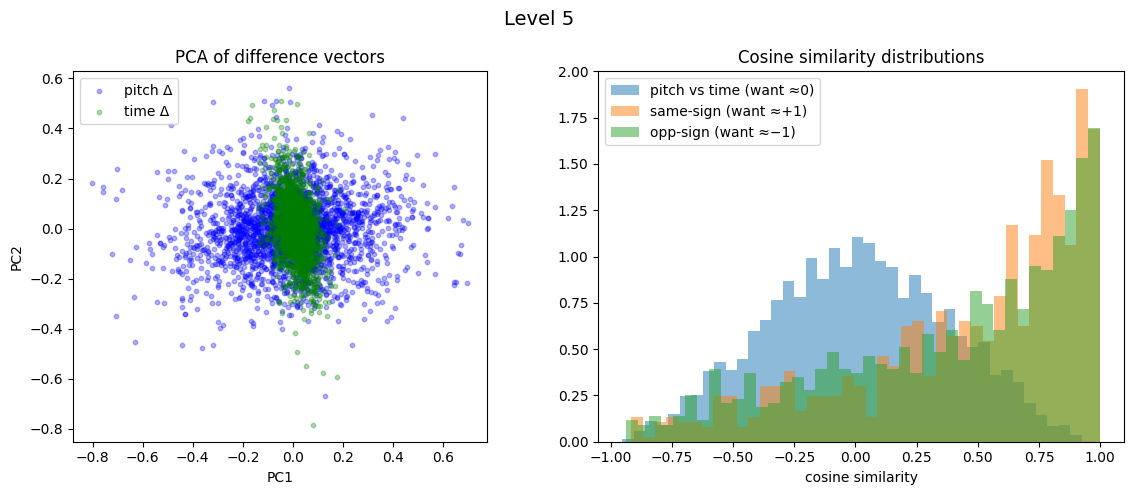
Level 5: cross=0.008, parallel=0.464, anti=0.389